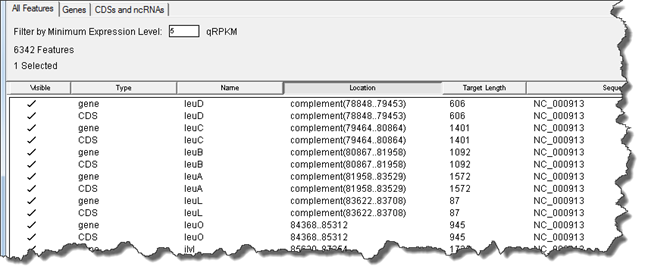
The Feature Tables allow you to view and manage all of the features within your current project. There are several different Feature Tables depending on the active selection and view:
|
Table Name |
Description |
To see the table, go to this view before choosing Features > Show Feature Table |
|
Features from [contig_name] |
Shows the features on both reference and constituent sequences. |
Alignment View (Contig > Alignment View) |
|
Features from [sequence_name] |
Shows the features only on the selected sequence. |
Alignment View (Contig > Alignment View) |
|
Features from Consensus of [contig_name] |
Shows the features on the consensus only. These are completely independent of the contig features. If you use Net Search > BLAST and select “Collect Features,” those features will be stored in this version of the table. |
Strategy View (Contig > Strategy View) |
Note that a separate Feature Table will open for each selected contig. If you select “Unlocated Contigs,” individual tables will open for each member of the group. You may wish to avoid selecting “Unlocated Contigs” if you are working with a large number of such contigs (as with many transcriptome or multi-chromosome genome assemblies).
The Feature Table window consists of three tabs that contain increasingly specific tables of features: All Features, Genes (only) and the subset of feature types chosen on the Input Reference Sequences page of the wizard (e.g. CDSs and ncRNAs only). Click on any tab to see a table for the features in that category.

The following tasks can be done within the Feature Tables:
|
Task |
How to… |
|
Display, hide, rearrange or sort data columns |
See Sorting and Rearranging Tables. For a list of available data columns, see Available Feature Table Columns. |
|
Optimize expression level filtering |
If desired, change the Filter by Minimum Expression Level to a custom value. Units are user-specified in the wizard (e.g., RPKM for RNA-Seq projects; weighted RPKM-CN for CNV projects). The default value is 5. |
|
View a selected feature in the Alignment or Strategy view |
As you select features in a Feature Table, the range of the feature is highlighted in the Alignment View, for features on constituent sequences, or the Strategy View, for features on the consensus sequence. |
|
Copy the sequence for a selected feature to the Unassembled Sequences window |
Sequences of features selected in a Feature Table may be added via drag and drop to the Unassembled Sequences window for additional assemblies. If the feature sequences you wish to assemble are located on a consensus sequence, you must first send the consensus sequence to another Lasergene application by using File > Send Consensus To before this option is enabled. |
|
Send the sequence of a selected feature to another Lasergene application |
Send the sequences of selected features to other Lasergene applications by using the File > Send Sequence Of Feature To command. |
|
Export a Feature Table |
|
|
Use a context menu to access editing commands |