Nephropath™ incorporates DNASTAR pipeline for validating processes against NIST “gold standard.”
The resources provided by the National Institute of Standards and Technology (NIST) Genome in a Bottle (GIAB) consortium promise to greatly improve the reliability of genetic assays. With these tools, laboratories can integrate performance measures directly within the workflow of their testing operations.
Nephropathology Associates, Inc. (Nephropath™), a leading U.S. laboratory in the interpretation of kidney biopsies, was motivated to use the NIST materials by the need to demonstrate proficiency in their NGS platform for purposes of CAP/CLIA certification. They were encouraged to look into GIAB by a representative from Illumina and, after a discussion with Justin Zook at the 2013 ASHG conference, decided that using the NIST data was the best option for them. The approach was also appealing because it would provide a measure of the lab’s accuracy as they would be able to compare their data with that of others who use the same controls.
As part of a collaborative project between Nephropath and DNASTAR, a new workflow has been added to DNASTAR’s assembly and variant calling software that supports use of the GIAB call sets.
The workflow is designed to work with a “gold standard” control of the user’s choice, such as the set of reference materials for the HapMap/1000 Genomes CEU female NA12878 developed by the GIAB consortium, as shown in Figure 1.
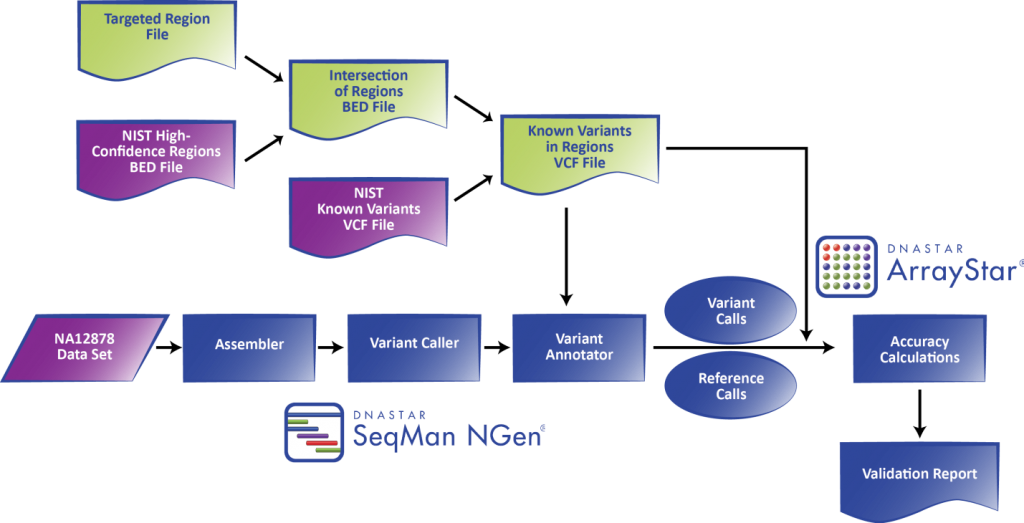
Figure 1. DNASTAR’s integrated Validated SNP Caller workflow used with NIST GIAB gold standard reference materials.
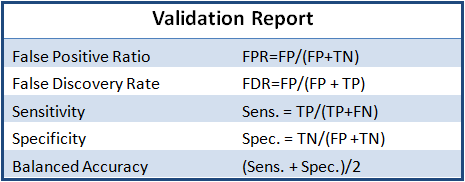
The purpose is to validate the efficacy of a procedure from sample prep through sequence analysis. At the end of the workflow, the lab obtains an automatically generated statistical report detailing the assembly sensitivity, specificity, and accuracy calculated according the ratios described in Table 1.
Nephropath is currently using the Illumina MiSeq and Agilent SureSelectQXT with custom probes for 301 genes involved in kidney disease. They use DNA from NA12878 purchased from Coriell Institute as a sequencing control on every run. Each run is a pool of 9 samples plus the control sequenced with the paired-end MiSeq® Reagent Kit v3 (150 cycle). The NA12878 control FASTQ files generated after the run are loaded into DNASTAR’s SeqMan NGen® software for mapping/alignment against the human genome reference sequence and variant calling using the “Templated assemblies with control” option. To delimit the regions of the genome used for validation, Nephropath uses a BED file of either their entire targeted region or one containing an intersection between the GIAB high quality regions and the targeted regions. The latter is preferred when the most accurate statistics are required. In this way, the NA12878 variant call set VCF file gets subsetted down to just the targeted regions using whichever BED file is selected. After the assembly is complete, every position specified by the BED file, including both variant and reference calls, is checked against the subsetted control VCF to determine true/false positives/negatives. Based on these annotated variant and reference call sets a validation report is generated by the DNASTAR ArrayStar® application, providing various statistics achieved at different sequencing depths and probability thresholds. An excerpt of such a report is given in Figure 2.

Figure 2. Nephropathology Associate’s Kidney Disease Gene Panel: Excerpts from a NA12878 Validation Report. Data provided by Marjorie Beggs (Nephropathology Associates) includes 301 genes from 13 renal disease categories.
The pipeline, along with early results, were presented at the recent GIAB consortium workshop in a roundup of emblematic case studies on using the GIAB materials.
Nephropath, in collaboration with DNASTAR, was recently awarded an SBIR phase I grant to further develop this workflow and software for clinical use. The long-term goal is to implement a fast, accurate and integrated workflow for clinical NGS.
For more information on the new validation workflow, download , “SeqMan NGen is a High Accuracy NGS Assembler: Assessment with NA12878 Reference Materials,” from the DNASTAR website.


Leave a Reply
Your email is safe with us.