For any function that allows you to choose a sequence, you can also choose to set ends for the sequence. There are four ways to choose a sub-range for a sequence in the Visual view:
- Via the Modify Sequence template.
- From a row with a Choose Sequence(s) button, right-click near the button and choose Add Sub-Range.
- From a row with a Choose Sequence(s) button, click on the plus icon (
) to the right of the row, then choose Add Sub-Range.
- From the Sequences section, apply Add Sub-Range by clicking on it or by drag-and-dropping it on the desired location in the Visual view.
The sequence range text boxes are initially populated with the terms “Left end” and “Right end.”
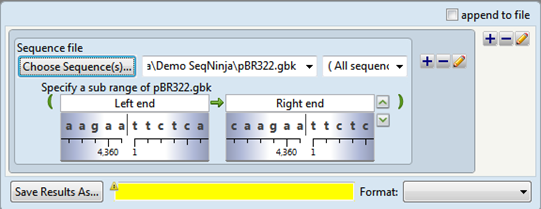
The following table shows tasks that can be done from within this dialog, or that affect its appearance.
| Task | How To |
|---|---|
| To reveal/hide the sequence, ruler and any features that are present; and to set ends based on features or by direct selection | Click the up/down arrows to the right of the range text boxes. See Work with Features for detailed instructions on how to view and work with sequence features. |
| To specify the desired range | Use the “Left end” and “Right end” boxes. You may type in a position or search for strings such as “atgc”. Alternatively, drag the “wheel” to go to a particular position number. Click any base to center the wheel on that base. You can reference the “left” of a sequence in the right-hand box, if desired (e.g., “left + 12”). |
| To skip to the left or right ends of the sequence | Click in a range text box, then use the left/right arrows that appear within the box. |
| To view feature information | Hover above a feature (if any are present) to see information about it, such as its left and right coordinates. |
| To reverse complement the current sub-range | Click the green arrow between the two range text boxes. |
IUPAC ambiguity codes are recognized both in the sequence and in the range boxes. For example, typing AAS (where S = C or G) into a range box would cause SeqNinja to look for the first instance of AAC or AAG in the sequence. Conversely, typing AAC into a range box would cause SeqNinja to look for the first instance of AAC, as well as any combination of bases and ambiguity codes that would allow for AAC (e.g., AAS, WWM, etc.).
Need more help with this?
Contact DNASTAR



 ) to the right of the row, then choose Add Sub-Range.
) to the right of the row, then choose Add Sub-Range.