The Strip Metadata function, located in the Functions section of the Toolkit panel, adds a metadata removal function to an existing script. This function may be used to remove specified annotations from a sequence.
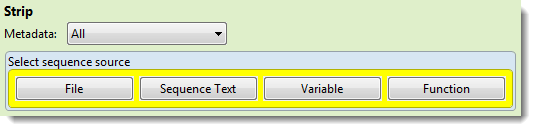
- Use the Metadata drop-down menu to choose which annotations to remove: All, Header (to leave features and remove only the header text), Features (to leave the header and remove all features), Features matching or Features except. If you choose one of the latter two options, a new row named “Features” opens just under the drop-down menu.

Use the drop-down menu to select the feature type to match (or not match): gene, CDS, exon, intron, mRNA, tRNA, promoter or misc_binding. To specify more than one feature type, use the plus (+) button to add additional “Features” rows. If you want to further limit the search to features matching (or not matching) particular qualifiers, check the Filter box, then choose a qualifier (gene, /product, /locus_tag, /note or /db_xref) from the drop-down menu to its right. In the right-most textbox, enter the text that the qualifier must match or not match (e.g., /gene = thrL) in order for the feature to be removed in the output file. You may use wildcards in this box if you wish (e.g., /gene = thr*).
- The “Select sequence source” part of this dialog requires you to select a sequence source. See Write to Results File for information on using the four buttons: File, Sequence Text, Variable and Function.
Need more help with this?
Contact DNASTAR


