The Features panel, represented by a “feature” icon (
Opening the Features panel:
Do any of the following:
- Click on the Features tab
- Use View > Features > Features .
- Press Ctrl+Alt+F (Windows) or Option+Cmd+F (Macintosh)

Using the Features panel:
| Task | How To |
|---|---|
| To create a new feature | 1) In any Protean 3D view, select the desired range for the feature. The selection can be discontinuous, but must be on a single molecule; it cannot span molecules. If you inadvertently create a feature from selections on multiple chains, multiple features with the same type and name will be added to the table. 2) Use the Create feature tool (  ). Clicking directly on the tool image adds a new “Region” feature. To create a feature of a different type, click on the arrow to the right of the tool and make a selection from the drop-down menu. ). Clicking directly on the tool image adds a new “Region” feature. To create a feature of a different type, click on the arrow to the right of the tool and make a selection from the drop-down menu.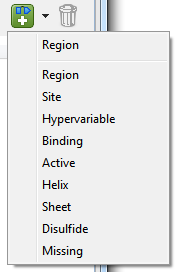 A feature named New feature 1 is created and is displayed graphically in the Analysis view. (User-created features are never visible in the Sequence view.) Subsequent features created in the same way are numbered incrementally. 3) If desired, edit the feature in the (feature) Options area. |
| To delete a feature | To delete a feature that was created using the above method, select it from the list in the Features panel and press the Delete feature tool ( ) or use the Features > Delete Feature menu command. Features that were imported with the sequence or structure cannot be deleted. ) or use the Features > Delete Feature menu command. Features that were imported with the sequence or structure cannot be deleted. |
| To order the tree alphabetically by feature type | Choose the Type radio button.Upon expanding a feature type, the features are sorted alphabetically by chain or sequence, and subsequently, by feature name. |
| To order features into their respective chains or sequences | Choose the Position radio button. Those features that traverse multiple chains are categorized as "Intermolecular." Upon expanding a chain or sequence, the features are organized by the order in which they appear in the chain or sequence. |
| To expand the feature list | Click the arrows to the left of items in the Names column. Expand arrows to the left of the check boxes can be used to reveal/hide additional elements. Expand arrow icons vary by operating system and may not be visible unless the mouse is hovering above them. Note that all structure files have two secondary structure assignments. The PDB secondary structure is the one annotated in the original PDB file. The KSD secondary structure is an automatic secondary structure assignment calculated by Protean 3D using the Kabsch and Sander DSSP method. See Work with Features for additional information. |
| To show/hide feature annotations in Sequence & Analysis views | Check/uncheck the box next to the feature ( ). Note that changing the visibility of any helix or sheet affects all helices and sheets in both views. ). Note that changing the visibility of any helix or sheet affects all helices and sheets in both views. |
The two areas related to the Features panel are described in separate topics:
Need more help with this?
Contact DNASTAR


