
Lasergene 17.3.1 is now available to download. This minor release includes new functionality and improvements for multiple sequence alignments in MegAlign Pro, and Sanger sequence assembly in SeqMan Ultra.
MegAlign Pro Enhancements and Updates
- Improved the guidance for choosing an alignment algorithm. MegAlign Pro now automatically chooses the best algorithm for your sequences based on the quantity, length, and the type of sequences you add. You still have the ability to change the alignment algorithm, if desired.
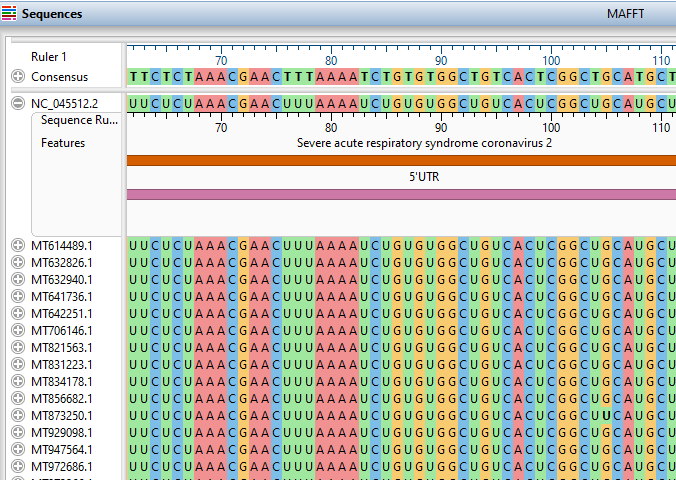
- Added the ability to specify a reference sequence for an alignment using the MAFFT algorithm, making it easy to quickly align thousands of sequences to a reference.
- Added the ability to search for disagreements between sequences.
- Expanded the ability to sort sequences in your project, including ordering sequences by name, length, distance, or other pairwise statistics.
- Added the ability to filter sequences by length, name, index, and content when adding sequences to your project, enabling you to select a smaller subset of a large set of sequences.
- Added some enhancements specifically for large phylogenetic trees, including the ability to zoom out farther, and the ability to collapse tree branches.
- Added the ability to flip individual branches in phylogenetic trees.
SeqMan Ultra Improvements and Updates
- Improved the guidance for Sanger multi-sample assemblies — we made it easier to find this popular option by adding a shortcut to the New Assembly window, which leads you through the appropriate steps for a multi-sample assembly.
- Added the Banana sequence sliding tool (originally from SeqMan Pro), which allows you to drag any Sanger sequence in the Alignment view to the left or right, as desired.
- Added the ability to reassemble contigs.





Leave a Reply
Your email is safe with us.