
Lasergene 17.3 is now available to download. This release includes significant additions for genomics research, specifically for long-read and viral genome assemblies, major improvements to MegAlign Pro, including the ability to quickly align thousands of viral genomes to known strains, and many enhancements and bug fixes for SeqMan Ultra and SeqBuilder Pro.
Lasergene Genomics Enhancements and Updates
- Expanded our viral genome assembly workflow to support PacBio and Oxford Nanopore long reads, including data from PCR-amplified fragments generated using the ARTIC Network protocols.
- Added support for PacBio Hifi long read data to our de novo genome assembly workflow, which produces a highly accurate assembly that can be edited in SeqMan Ultra.
- Laid the groundwork in GenVision Pro for significant changes to come, including adding a new track for multi-sample, whole genome navigation, an expanded Explorer panel for easily adding and removing assemblies, improved search functionality, and complete integration with SeqMan NGen.
- Addressed some issues under the hood for RNA-Seq and de novo transcriptome assemblies, improving success rates for these types of projects.
MegAlign Pro Enhancements and Updates
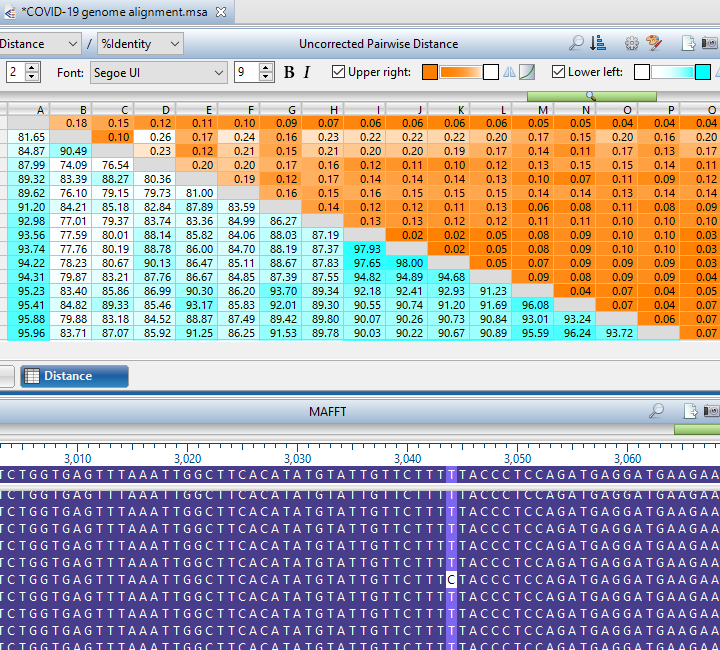
- Updated the MAFFT alignment algorithm in MegAlign Pro to the significantly faster MAFFT7, resulting in much quicker alignments for viral genomes and 16S data sets.
- Added support to easily import several widely-used alignment formats, including Vector NTI, GCG, Geneious, and MEGA files, as well as support for exporting MEGA files.
- Added an alignment report that can be configured and exported.
- Added alignment color schemes for hydrophobicity, percent identity, and others similar to ClustalX and UGene.
- Added the ability to color gaps, making it easier to highlight the differences between sequences.
- Made significant improvements to support large distance tables, including the addition of color gradients and zoom controls, making it easier to identify groups of interest in large data sets.
- Made general improvements to the distance table, including the ability to calculate % dissimilar, expanded toolbars for easier navigation and style application, and the ability to export images for publication or collaboration.
- Added the ability to sort sequences by distance and other measures.
- Added the ability to search the consensus or individual sequences for a specific string of nucleotides or amino acids.
SeqMan Ultra updates include:
- New support for importing Sequencher (.spf) files
As well as fixes for the following issues:
- An error when importing some sequence assembly files
- Difficulties with exporting reads
- Spurious coloring of non-variant bases
- Long read alignment track performance issues
- Incorrect reports for long read assemblies
- Difficulties with exporting images
- Sanger traces lost in SeqMan NGen assemblies
- Sanger secondary (heterozygous) peaks and deletions (“-“) not being identified as variants
- Variant table not updating after edits in Sanger assemblies
- Marked reference order of assembly issues in Sanger assemblies
SeqBuilder Pro updates include fixes for the following issues:
- Support for the current Geneious format
- “Find” function not working on macOS Big Sur
- Issues with adding custom vectors
- A crash when copying primer sequences
- SeqMan Ultra not opening in verify clone workflow
- A crash occurring during TA cloning
- Batch translate not working with custom parameters
If you are a current customer, upgrades are always included as part of your service plan. If your service plan has expired, please request a quote to renew!





Leave a Reply
Your email is safe with us.