Whether you are coming from Part A: Create a templated assembly in SeqMan NGen or the optional Part B: Filter for variants of interest in ArrayStar, the next step is to view the variants in SeqMan Pro.
- If you are coming from Part B, the sample is automatically selected and its Alignment view is opened. Continue directly to Step 2. If you are coming from Part A, launch SeqMan Pro and use File > Open to open the .assembly file.
- Choose Variant > Variant Report. (If desired variants are being filtered out, you may need to click on the Show All button.)
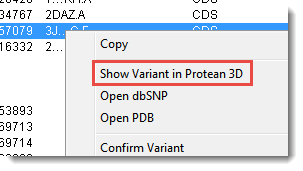
- Click on the PDB ID column header to sort items with PDB IDs to the top. Select one or more rows, then right-click within the selection and choose Show Variant in Protean 3D (or use Variant > Show Variant in Protean 3D).
Continue to Part D: View the protein structure in Protean 3D.
Need more help with this?
Contact DNASTAR


